Abstract
Objectives
To evaluate for the first time the performance of a deep learning method based on no-new-Net for fully automated segmentation and volumetric measurements of intracerebral hemorrhage (ICH), intraventricular extension of intracerebral hemorrhage (IVH), and perihematomal edema (PHE) in primary ICH on CT.
Methods
Three hundred and eighty primary ICH patients who underwent CT at hospital arrival were divided into a training cohort (n = 300) and a validation cohort (n = 80). An independent cohort with 80 patients was used for testing. Ground truth (segmentation masks) was manually generated by radiologists. Model performance on lesion segmentation and volumetric measurement of ICH, IVH, and PHE were evaluated by comparing the model results with the segmentations performed by radiologists.
Results
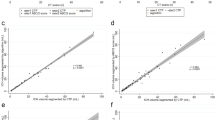
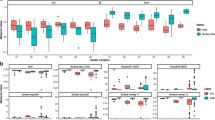
In the test cohort, the Dice scores of lesion segmentation were 0.92, 0.79, and 0.71 for ICH, IVH, and PHE, respectively. The sensitivities were 0.93 for ICH, 0.88 for IVH, and 0.81 for PHE. The positive predictive values were 0.92, 0.76, and 0.69 for ICH, IVH, and PHE, respectively. Excellent concordance (concordance correlation coefficients [CCCs] ≥ 0.98) of ICH and IVH and good concordance of PHE (CCCs ≥ 0.92) were demonstrated between manually and automatically measured volumes. The model took approximately 15 s to provide automatic segmentation and volume analysis for each patient.
Conclusion
Our model demonstrates good reliability for automatic segmentation and volume measurement of ICH, IVH, and PHE in primary ICH, which can be useful to reduce the effort and time of doctors to calculate volumes of ICH, IVH, and PHE.
Key Points
• Deep learning algorithms can provide automatic and reliable assessment of intracerebral hemorrhage, intraventricular hemorrhage, and perihematomal edema on CT.
• Non-contrast CT-based deep learning method can be helpful to provide efficient and accurate measurements of ICH, IVH, and PHE in primary ICH patients, thereby reducing the effort and time of doctors to segment and calculate volumes of ICH, IVH, and PHE in primary ICH patients.



Similar content being viewed by others
Abbreviations
- CCCs:
-
Concordance correlation coefficients
- GCS:
-
Glasgow Coma Scale
- ICH:
-
Intracerebral hemorrhage
- IVH:
-
Intraventricular extension of intracerebral hemorrhage
- PHE:
-
Perihematomal edema
- PPV:
-
Positive predictive value
References
Cordonnier C, Demchuk A, Ziai W, Anderson CS (2018) Intracerebral haemorrhage: current approaches to acute management. Lancet 392:1257–1268
Pinho J, Costa AS, Araújo JM, Amorim JM, Ferreira C (2019) Intracerebral hemorrhage outcome: a comprehensive update. J Neurol Sci 398:54–66
Hanley DF (2009) Intraventricular hemorrhage: severity factor and treatment target in spontaneous intracerebral hemorrhage. Stroke 40:1533–1538
Irvine H, Male S, Robertson J, Bell C, Bentho O, Streib C (2019) Reduced intracerebral hemorrhage and perihematomal edema volumes in diabetics on sulfonylureas. Stroke 50:995–998
Urday S, Kimberly WT, Beslow LA et al (2015) Targeting secondary injury in intracerebral haemorrhage--perihaematomal oedema. Nat Rev Neurol 11:111–122
Ironside N, Chen C-J, Ding D, Mayer SA, Connolly ES Jr (2019) Perihematomal edema after spontaneous intracerebral hemorrhage. Stroke 50:1626–1633
Yang J, Arima H, Wu G et al (2015) Prognostic significance of perihematomal edema in acute intracerebral hemorrhage: pooled analysis from the intensive blood pressure reduction in acute cerebral hemorrhage trial studies. Stroke 46:1009–1013
Dastur CK, Yu W (2017) Current management of spontaneous intracerebral haemorrhage. Stroke Vasc Neurol 2:21–29
Cheung RTF, Zou L-Y (2003) Use of the original, modified, or new intracerebral hemorrhage score to predict mortality and morbidity after intracerebral hemorrhage. Stroke 34:1717–1722
Ziai WC, Carhuapoma JR (2018) Intracerebral hemorrhage. Continuum (Minneap Minn) 24:1603–1622
Volbers B, Willfarth W, Kuramatsu JB et al (2016) Impact of perihemorrhagic edema on short-term outcome after intracerebral hemorrhage. Neurocrit Care 24:404–412
Appelboom G, Bruce SS, Hickman ZL et al (2013) Volume-dependent effect of perihaematomal oedema on outcome for spontaneous intracerebral haemorrhages. J Neurol Neurosurg Psychiatry 84:488–493
Urday S, Beslow LA, Dai F et al (2016) Rate of perihematomal edema expansion predicts outcome after intracerebral hemorrhage. Crit Care Med 44:790–797
Yeatts SD, Palesch YY, Moy CS, Selim M (2013) High dose deferoxamine in intracerebral hemorrhage (HI-DEF) trial: rationale, design, and methods. Neurocrit Care 19:257–266
Kollmar R, Staykov D, Dörfler A, Schellinger PD, Schwab S, Bardutzky J (2010) Hypothermia reduces perihemorrhagic edema after intracerebral hemorrhage. Stroke 41:1684–1689
Freeman WD, Barrett KM, Bestic JM, Meschia JF, Broderick DF, Brott TG (2008) Computer-assisted volumetric analysis compared with ABC/2 method for assessing warfarin-related intracranial hemorrhage volumes. Neurocrit Care 9:307–312
Huttner HB, Steiner T, Hartmann M et al (2006) Comparison of ABC/2 estimation technique to computer-assisted planimetric analysis in warfarin-related intracerebral parenchymal hemorrhage. Stroke 37:404–408
Chang K, Beers AL, Bai HX et al (2019) Automatic assessment of glioma burden: a deep learning algorithm for fully automated volumetric and bidimensional measurement. Neuro Oncol 21:1412–1422
Brugnara G, Isensee F, Neuberger U et al (2020) Automated volumetric assessment with artificial neural networks might enable a more accurate assessment of disease burden in patients with multiple sclerosis. Eur Radiol 30:2356–2364
Kickingereder P, Isensee F, Tursunova I et al (2019) Automated quantitative tumour response assessment of MRI in neuro-oncology with artificial neural networks: a multicentre, retrospective study. Lancet Oncol 20:728–740
Ye H, Gao F, Yin Y et al (2019) Precise diagnosis of intracranial hemorrhage and subtypes using a three-dimensional joint convolutional and recurrent neural network. Eur Radiol 29:6191–6201
Ronneberger O, Fischer P, Brox T (2015) U-net: convolutional networks for biomedical image segmentation. Available via https://arxiv.org/abs/1505.04597. Accessed 18 May 2015
Çiçek Ö, Abdulkadir A, Lienkamp SS, Brox T, Ronneberger O (2016) 3D U-Net: learning dense volumetric segmentation from sparse annotation. Available via https://arxiv.org/abs/1606.06650. Accessed 21 Jun 2016
Milletari F, Navab N, Ahmadi S-A (2016) V-net: fully convolutional neural networks for volumetric medical image segmentation. IEEE. Available via https://arxiv.org/abs/1606.04797. Accessed 15 Jun 2016
Casamitjana A, Catà M, Sánchez I, Combalia M, Vilaplana V (2017) Cascaded V-Net using ROI masks for brain tumor segmentation. Available via https://arxiv.org/abs/1812.11588. Accessed 30 Dec 2018
Isensee F, Petersen J, Klein A et al (2018) nnu-net: self-adapting framework for u-net-based medical image segmentation. Available via https://arxiv.org/abs/1809.10486. Accessed 27 Sep 2018
Li X, Chen H, Qi X, Dou Q, Fu C-W, Heng P-A (2018) H-DenseUNet: hybrid densely connected UNet for liver and tumor segmentation from CT volumes. IEEE Trans Med Imaging 37:2663–2674
Volbers B, Staykov D, Wagner I et al (2011) Semi-automatic volumetric assessment of perihemorrhagic edema with computed tomography. Eur J Neurol 18:1323–1328
Isensee F, Jäger PF, Kohl SAA, Petersen J, Maier-Hein KH (2019) Automated design of deep learning methods for biomedical image segmentation. Available via https://arxiv.org/abs/1904.08128. Accessed 17 Apr 2019
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. Available via https://arxiv.org/abs/1412.6980. Accessed 22 Dec 2014
Drozdzal M, Vorontsov E, Chartrand G, Kadoury S, Pal C (2016) The importance of skip connections in biomedical image segmentation. Available via https://arxiv.org/abs/1608.04117. Accessed 14 Aug 2016
Chen L-C, Papandreou G, Kokkinos I, Murphy K, Yuille AL (2017) DeepLab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected CRFs. Available via https://arxiv.org/abs/1606.00915. Accessed 2 Jun 2016
Zhao X, Wu Y, Song G, Li Z, Zhang Y, Fan Y (2018) A deep learning model integrating FCNNs and CRFs for brain tumor segmentation. Med Image Anal 43:98–111
Patel A, Schreuder F, Klijn CJM et al (2019) Intracerebral haemorrhage segmentation in non-contrast CT. Sci Rep 9:17858
Ironside N, Chen CJ, Mutasa S et al (2019) Fully automated segmentation algorithm for hematoma volumetric analysis in spontaneous intracerebral hemorrhage. Stroke 50:3416–3423
Dhar R, Falcone GJ, Chen Y et al (2020) Deep learning for automated measurement of hemorrhage and perihematomal edema in supratentorial intracerebral hemorrhage. Stroke 51:648–651
Chang PD, Kuoy E, Grinband J et al (2018) Hybrid 3D/2D convolutional neural network for hemorrhage evaluation on head CT. AJNR Am J Neuroradiol 39:1609–1616
Davis SM, Broderick J, Hennerici M et al (2006) Hematoma growth is a determinant of mortality and poor outcome after intracerebral hemorrhage. Neurology 66:1175–1181
Al-Shahi Salman R, Frantzias J, Lee RJ et al (2018) Absolute risk and predictors of the growth of acute spontaneous intracerebral haemorrhage: a systematic review and meta-analysis of individual patient data. Lancet Neurol 17:885–894
Morotti A, Boulouis G, Dowlatshahi D et al (2019) Standards for detecting, interpreting, and reporting noncontrast computed tomographic markers of intracerebral hemorrhage expansion. Ann Neurol 86:480–492
Hallevi H, Dar NS, Barreto AD et al (2009) The IVH score: a novel tool for estimating intraventricular hemorrhage volume: clinical and research implications. Crit Care Med 37:969–974 e961
Chan E, Anderson CS, Wang X et al (2015) Significance of intraventricular hemorrhage in acute intracerebral hemorrhage: intensive blood pressure reduction in acute cerebral hemorrhage trial results. Stroke 46:653–658
Ironside N, Chen CJ, Mutasa S et al (2020) Fully automated segmentation algorithm for perihematomal edema volumetry after spontaneous intracerebral hemorrhage. Stroke 51:815–823
Lin L, Dou Q, Jin YM et al (2019) Deep learning for automated contouring of primary tumor volumes by MRI for nasopharyngeal carcinoma. Radiology 291:677–686
Rudie JD, Weiss DA, Saluja R et al (2019) Multi-disease segmentation of gliomas and white matter hyperintensities in the BraTS data using a 3D convolutional neural network. Front Comput Neurosci 13:84
Egger C, Opfer R, Wang C et al (2017) MRI FLAIR lesion segmentation in multiple sclerosis: does automated segmentation hold up with manual annotation? Neuroimage Clin 13:264–270
Castro DC, Walker I, Glocker B (2020) Causality matters in medical imaging. Nat Commun 11:3673
Funding
This study has received funding from Shanghai Municipal Health Commission (grant number 2019SY061).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Zhenwei Yao.
Conflict of interest
The authors of this manuscript declare relationships with the following company: Ping An Technology (Shenzhen) Co., Ltd.
Statistics and biometry
One of the authors has significant statistical expertise.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 466 kb)
Rights and permissions
About this article
Cite this article
Zhao, X., Chen, K., Wu, G. et al. Deep learning shows good reliability for automatic segmentation and volume measurement of brain hemorrhage, intraventricular extension, and peripheral edema. Eur Radiol 31, 5012–5020 (2021). https://doi.org/10.1007/s00330-020-07558-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-020-07558-2